Metagenomic Analysis
made Free
The no-code, cloud-based bioinformatics tool for researchers
112+
121+
320,000+

Fast Pipelines, No-Code Platform
Metagenomic Pipeline

Characterize microbes present in your samples, including viruses, bacteria, and eukaryotes
Sequencing Platform:
Illumina & Nanopore
Data Types Supported:
Shotgun (or random) data
Illumina Average Runtime:
3 hours
Nanopore Average Runtime:
26 mins
Antimicrobial Resistance Pipeline

Detect antimicrobial resistant genes in your samples
Sequencing Platform:
Illumina
Data Types Supported:
Shotgun (or random) and whole genome sequence data
Illumina Average Runtime:
16 mins
Viral Consensus Genome Pipeline

Create consensus genomes for any virus
Sequencing Platform:
Illumina
Data Types Supported:
Target enrichment (e.g., MSSPE), PCR, whole genome, and metagenomic sequencing data
Illumina Average Runtime:
18 mins



Get Started with 4 Simple Steps
All you need is a laptop and an internet connection to analyze your data.
Upload Samples
We accept raw sequencing data from Illumina and Nanopore
Run Pipeline
Samples run concurrently in the cloud through our automated pipeline
View Report
Our report page provides insights and key metrics necessary for your analysis
Visualize Data
Create heatmaps and quality control charts to help draw conclusions across samples

Engineered for Scientists
CZ ID automates trusted pipelines, so that you can focus your time on analysis and next steps.
Fully Automated
Detailed analyses, no coding required
Fast and Scalable
Rapid, concurrent sample analysis right from your laptop
Private and Secure
We protect, but never sell or own your data
Powerful, Customizable Data Visualizations



Heatmap
Customize the heatmap to answer your research questions
Coverage Visualization
Check out the coverage breadth and depth for taxa of interest
Taxonomic Tree
View a phylogram of your sample’s microbes
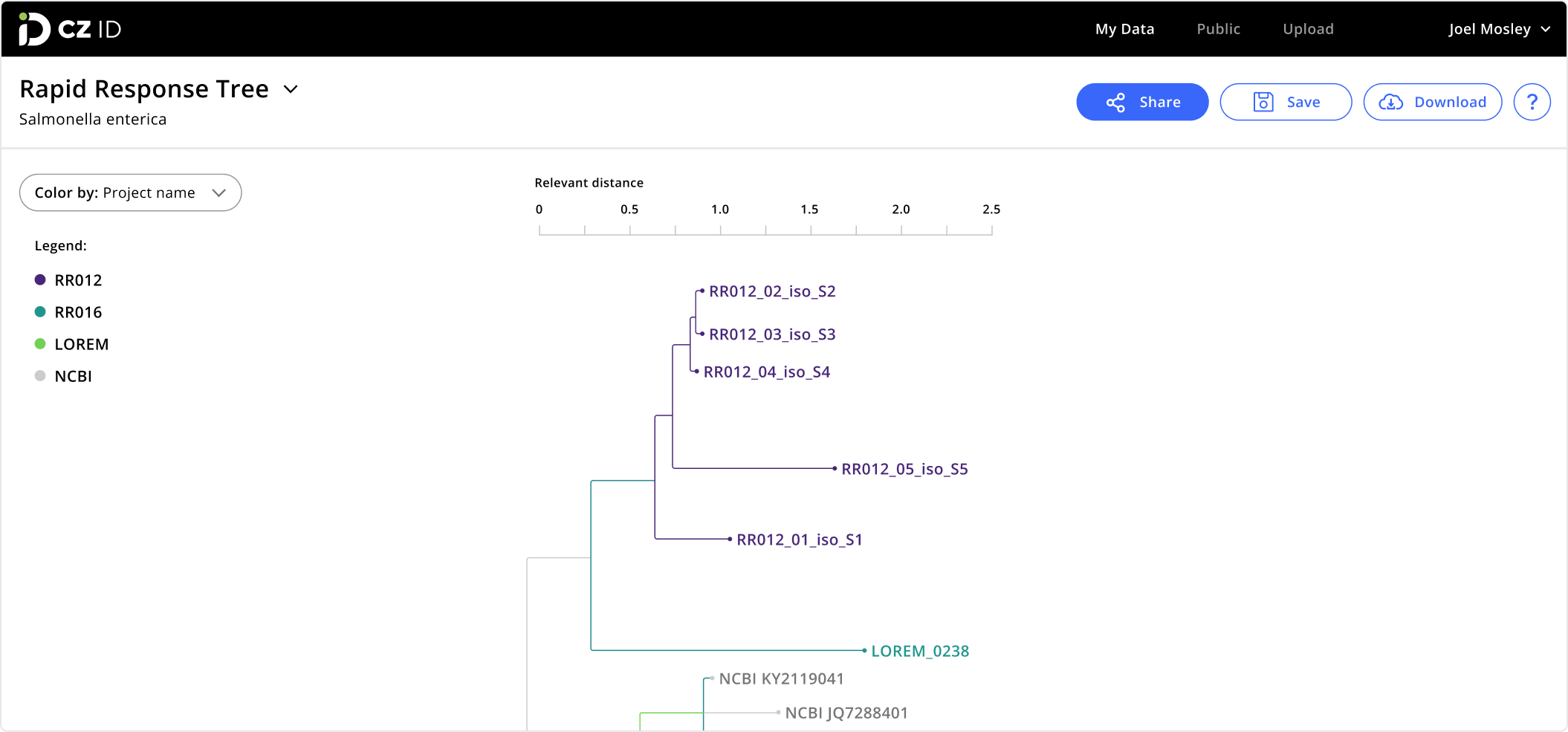
Phylogenetic Tree
Evaluate sequence similarity to identify potential outbreaks
Heatmap
Customize the heatmap to answer research questions

Coverage Visualization
Check out the coverage breadth and depth for taxon hits

Taxonomic Tree
View a phylogram of your samples microbes

Phylogenetic Tree
Evaluate sequence similarity to identify potential outbreaks

We Work Closely with Trusted Partners
CZ ID collaborates with trusted partners in the bioinformatics, infectious disease, and sequencing ecosystem.




Check out Our Paper in GigaScience
We describe CZ ID (formerly IDseq), its capabilities, and how the tool was validated. CZ ID is a continuously evolving service. For the most up to date analysis pipeline, check out our Github page.
See how RESEARCHERS are using CZ ID
Ramachandran, P. S., et al. “Integrating central nervous system metagenomics and host response for diagnosis of tuberculosis meningitis and its mimics.” Nature communications 13.1 (2022): 1675.
View Paper
Saha, Senjuti, et al. “Unbiased metagenomic sequencing for pediatric meningitis in Bangladesh reveals neuroinvasive chikungunya virus outbreak and other unrealized pathogens.” MBio 10.6 (2019): 10-1128.
View Paper
Baillie, Vicky L., et al. “Metagenomic sequencing of post-mortem tissue samples for the identification of pathogens associated with neonatal deaths.” Nature Communications 14.1 (2023): 5373.
View Paper
Bohl, Jennifer A., et al. “Discovering disease-causing pathogens in resource-scarce Southeast Asia using a global metagenomic pathogen monitoring system.” Proceedings of the National Academy of Sciences 119.11 (2022): e2115285119.
View Paper
Frequently Asked Questions
There is no limit on the amount of data that you can upload to CZ ID.
Yes, CZ ID is committed to remaining a free tool.
Raw sample data (genetic sequence files (ex: FASTA/FASTQ)) is not shared with any other CZ ID user, nor is it ever accessed by anyone working on CZ ID unless specifically requested by a user, such as to debug an issue. Read more in CZ ID’s Privacy Policy.
Upon upload, raw sample data (genetic sequence files (ex: FASTA/FASTQ)) is processed through our data pipeline and all host (ex: human, mosquito) genetic information is filtered out. We always filter out all human genetic information, regardless of host. Read more in CZ ID’s Privacy Policy.
Once created, your account will be maintained in accordance with our Terms of Use. You can request to delete your account at any time by contacting our team.